Gene
Gene:
CTAGE5 (official gene symbol)Other symbol:
CTAGE5CT family:
CT21CT identifier:
CT21.3Aliases from NCBI:
MEA6 , MGEA , MGEA11 , MGEA6RefSeq
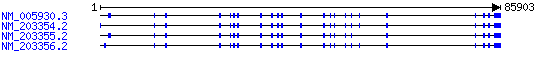
Nucleotide : NM_203356.2 | NM_203355.2 | NM_203354.2 | NM_005930.3
Phylogenetic relationships with CT genes
This gene belongs to the CTAGE family.
Gene structure and chromosomal localization
CTAGE5 was mapped to 14q13.3 by somatic hybrid panel mapping and fluorescence in situ hybridization (Heckel et al., 1997 PMID: 9356211).
Chromosome band: 14q13.3
See this gene in the UCSC Genome Browser :
See this gene in the NCBI Map Viewer Genome Browser :
SNPs
Show| ACCESSION | ALLELES | HET | SE_HET | FXN_CLASS | RESIDUE | AA_POSITION | VALIDATED |
|---|---|---|---|---|---|---|---|
| rs1060878 | A/G | 0.137487 | 0.22325 | missense | R | 708 | YES |
| rs1060878 | A/G | 0.137487 | 0.22325 | reference | G | 708 | YES |
| rs1060878 | A/G | 0.137487 | 0.22325 | missense | R | 694 | YES |
| rs1060878 | A/G | 0.137487 | 0.22325 | reference | G | 694 | YES |
| rs1060878 | A/G | 0.137487 | 0.22325 | missense | R | 725 | YES |
| rs1060878 | A/G | 0.137487 | 0.22325 | reference | G | 725 | YES |
| rs1060878 | A/G | 0.137487 | 0.22325 | missense | R | 737 | YES |
| rs1060878 | A/G | 0.137487 | 0.22325 | reference | G | 737 | YES |
| rs1140952 | A/G | 0.382071 | 0.212267 | reference | I | 655 | YES |
| rs1140952 | A/G | 0.382071 | 0.212267 | missense | V | 655 | YES |
| rs1140952 | A/G | 0.382071 | 0.212267 | reference | I | 669 | YES |
| rs1140952 | A/G | 0.382071 | 0.212267 | missense | V | 669 | YES |
| rs1140952 | A/G | 0.382071 | 0.212267 | reference | I | 698 | YES |
| rs1140952 | A/G | 0.382071 | 0.212267 | missense | V | 698 | YES |
| rs1140952 | A/G | 0.382071 | 0.212267 | reference | I | 686 | YES |
| rs1140952 | A/G | 0.382071 | 0.212267 | missense | V | 686 | YES |
| rs1140952 | A/G | 0.382071 | 0.212267 | missense | I | 655 | YES |
| rs1140952 | A/G | 0.382071 | 0.212267 | reference | V | 655 | YES |
| rs1140952 | A/G | 0.382071 | 0.212267 | missense | I | 669 | YES |
| rs1140952 | A/G | 0.382071 | 0.212267 | reference | V | 669 | YES |
| rs1140952 | A/G | 0.382071 | 0.212267 | missense | I | 698 | YES |
| rs1140952 | A/G | 0.382071 | 0.212267 | reference | V | 698 | YES |
| rs1140952 | A/G | 0.382071 | 0.212267 | missense | I | 686 | YES |
| rs1140952 | A/G | 0.382071 | 0.212267 | reference | V | 686 | YES |
| rs1950952 | C/G | 0.499 | 0.0223418 | missense | Q | 359 | YES |
| rs1950952 | C/G | 0.499 | 0.0223418 | reference | E | 359 | YES |
| rs1950952 | C/G | 0.499 | 0.0223418 | missense | Q | 330 | YES |
| rs1950952 | C/G | 0.499 | 0.0223418 | reference | E | 330 | YES |
| rs1950952 | C/G | 0.499 | 0.0223418 | missense | Q | 347 | YES |
| rs1950952 | C/G | 0.499 | 0.0223418 | reference | E | 347 | YES |
| rs1950952 | C/G | 0.499 | 0.0223418 | reference | Q | 359 | YES |
| rs1950952 | C/G | 0.499 | 0.0223418 | missense | E | 359 | YES |
| rs1950952 | C/G | 0.499 | 0.0223418 | reference | Q | 330 | YES |
| rs1950952 | C/G | 0.499 | 0.0223418 | missense | E | 330 | YES |
| rs1950952 | C/G | 0.499 | 0.0223418 | reference | Q | 347 | YES |
| rs1950952 | C/G | 0.499 | 0.0223418 | missense | E | 347 | YES |
| rs7140561 | C/T | 0.056404 | 0.158179 | reference | A | 5 | YES |
| rs7140561 | C/T | 0.056404 | 0.158179 | missense | V | 5 | YES |
| rs7140561 | C/T | 0.056404 | 0.158179 | missense | A | 5 | YES |
| rs7140561 | C/T | 0.056404 | 0.158179 | reference | V | 5 | YES |
| rs10162564 | A/G | 0.0281024 | 0.115158 | reference | K | 249 | YES |
| rs10162564 | A/G | 0.0281024 | 0.115158 | missense | E | 249 | YES |
| rs10162564 | A/G | 0.0281024 | 0.115158 | reference | K | 237 | YES |
| rs10162564 | A/G | 0.0281024 | 0.115158 | missense | E | 237 | YES |
| rs10162564 | A/G | 0.0281024 | 0.115158 | reference | K | 220 | YES |
| rs10162564 | A/G | 0.0281024 | 0.115158 | missense | E | 220 | YES |
| rs17109109 | A/G | 0.0285654 | 0.116046 | reference | N | 374 | YES |
| rs17109109 | A/G | 0.0285654 | 0.116046 | missense | S | 374 | YES |
| rs17109109 | A/G | 0.0285654 | 0.116046 | reference | N | 362 | YES |
| rs17109109 | A/G | 0.0285654 | 0.116046 | missense | S | 362 | YES |
| rs17109109 | A/G | 0.0285654 | 0.116046 | reference | N | 345 | YES |
| rs17109109 | A/G | 0.0285654 | 0.116046 | missense | S | 345 | YES |
| rs34761053 | G/T | 0.0259695 | 0.110952 | reference | K | 213 | NO |
| rs34761053 | G/T | 0.0259695 | 0.110952 | missense | Q | 213 | NO |
| rs34761053 | G/T | 0.0259695 | 0.110952 | reference | K | 242 | NO |
| rs34761053 | G/T | 0.0259695 | 0.110952 | missense | Q | 242 | NO |
| rs34761053 | G/T | 0.0259695 | 0.110952 | reference | K | 230 | NO |
| rs34761053 | G/T | 0.0259695 | 0.110952 | missense | Q | 230 | NO |
| rs34909359 | A/C | 0.0587695 | 0.161031 | reference | P | 9 | YES |
| rs34909359 | A/C | 0.0587695 | 0.161031 | coding-synonymous | P | 9 | YES |
| rs35626166 | C/G | 0.0259695 | 0.110952 | missense | L | 221 | NO |
| rs35626166 | C/G | 0.0259695 | 0.110952 | reference | V | 221 | NO |
| rs35626166 | C/G | 0.0259695 | 0.110952 | missense | L | 250 | NO |
| rs35626166 | C/G | 0.0259695 | 0.110952 | reference | V | 250 | NO |
| rs35626166 | C/G | 0.0259695 | 0.110952 | missense | L | 238 | NO |
| rs35626166 | C/G | 0.0259695 | 0.110952 | reference | V | 238 | NO |
| rs36060072 | C/G | 0.0259695 | 0.110952 | reference | P | 491 | NO |
| rs36060072 | C/G | 0.0259695 | 0.110952 | missense | A | 491 | NO |
| rs36060072 | C/G | 0.0259695 | 0.110952 | reference | P | 520 | NO |
| rs36060072 | C/G | 0.0259695 | 0.110952 | missense | A | 520 | NO |
| rs36060072 | C/G | 0.0259695 | 0.110952 | reference | P | 508 | NO |
| rs36060072 | C/G | 0.0259695 | 0.110952 | missense | A | 508 | NO |